Text-mining exercises
Learning objectives
In these exercises, we will use a variety of text-mining tools and databases based on text mining to interpret the associations of genes and diseases. The exercises will teach you how to:
- automatically highlight named entities in a web page
- use named entity recognition for synonym-aware information retrieval
- extract associations based on cooccurrence of entities in the literature
Prerequisites
All exercises are purely web-based.
Exercise 1
In this exercise we will first introduce the basics of text mining: 1) dictionary-based named entity recognition and 2) how this can used to help retrieve literature. Afterwards we will move on to how one can use the complete literature to 3) extract associations between entities and finally 4) how these associations can be used for knowledge discovery.
1.1 Named Entity Recognition
The goal of named entity recognition (NER) is to find names mentioned in text and resolve them to the underlying biomedical entities (document → entity A, entity B, entity C). To illustrate this, we will use the EXTRACT tool, which is designed to use NER to support manual database curation.
Install the EXTRACT bookmarklet as described on the EXTRACT website.
Hint: If the bookmarks toolbar is not showing in Firefox then go the File menu bar and select View → Toolbars → Bookmarks Toolbar → Always show. If it is not showing on Chrome go to the menu bar and select View → Always show bookmarks toolbar
Open the paper “Age-Dependent Brain Gene Expression and Copy Number Anomalies in Autism Suggest Distinct Pathological Processes at Young Versus Mature Ages” (Chow et al., 2012) and click the EXTRACT bookmarklet. After a short time, terms should be highlighted in the text.
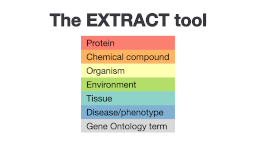
1. What do the different colors mean? How many different types of biomedical entities can you see in the abstract?
By clicking or hovering over a tagged term, you will get a popup that includes its standard name, entity type, database or ontology identifier, and a link to its reference record. Move to the Results section of the paper and click or hover over autism and NFBD1.
2. What is the main name for autism in Disease Ontology? What is the difference between NFBD1 and MDC1?
3. What is the Disease ontology identifier for autism and what is the ENSEMBL identifier for NFBD1?
Select the Author Summary (the paragraph, not the actual words) in the paper and click the EXTRACT bookmarklet. Hover over terms in the text or lines in the table.
4. What happens when you hover over the terms under the selected text section in the window that pops up?
Use the buttons in the bottom of the popup to copy the data into a spreadsheet (e.g. Microsoft excel/Google sheets) or text file (e.g. Notepad/Notepad++) or save it in tabular (tsv/csv) format.
5. Which information is then provided in addition to what is shown in the popup?
1.2 Information retrieval
The goal of information retrieval (IR) is to find the documents pertaining to a topic of interest. When the topic is a biological entity (A), NER can be used to index the literature and thereby support retrieval of relevant documents (A → documents).
We run the same NER system used in EXTRACT on entire PubMed every week and make the results available through a suite of web resources. One such resource is DISEASES. While primarily intended to view disease–gene associations extracted from literature, it can also be used for information retrieval.
Click the following link to retrieve abstracts that mention the gene SCN2A:
https://diseases.jensenlab.org/Entity?documents=10&type1=9606&id1=ENSP00000490107
6. Do the abstracts shown in the first two pages all highlight SCN2A or are there other words highlighted? If the latter is true, can you think of a reason why?
You can similarly use NER to retrieve abstracts for any disease in the Disease Ontology. For example, the following query will retrieve abstracts for neurodegenerative disease (DOID:1289):
https://diseases.jensenlab.org/Entity?documents=10&type1=-26&id1=DOID:1289
7. Which diseases are highlighted in the abstracts? Can you think of the reason why other diseases are also mentioned?
1.3 Relation extraction
The goal of cooccurrence-based relation extraction (RE) is to link entities (A, B) to each other based on them being mentioned together in documents (A → documents → B).
Go to https://diseases.jensenlab.org/ and query for LRRK. You are now presented with two options, since there are two gene names starting with LRRK, namely LRRK1 and LRRK2. Click on the LRRK2 row to view the disease associations for this gene.
8. Which disease is most strongly associated with LRKK2 according to text mining?
Click on Parkinson's disease in the text-mining table.
9. Do the abstracts in fact support an association between Parkinson's disease and LRRK2?
Now click on Alzheimer's disease.
10. Do the abstracts support an association in this case too? If not can you think of reasons why this association has such a high score?
Cooccurrence-based relation extraction is a very generic approach, which can be used to find associations between any two types of entities for which we can do NER. For example, we can use the same approach to extract human phenotypes associated with SCN1B from the mammalian phenotypes database:
https://phenotypes.jensenlab.org/Entity?textmining=20&type1=9606&type2=-36&id1=ENSP00000396915
Click on Seizures and look at the associations between that term and SCN1B.
11. Is this association supported by only few papers or is it well established in the literature? Do all the papers that mention the two terms actually support this association?
Note: You do not need to provide the actual number of papers supporting the association.
Exercise 2
In this exercise, we will focus on how one can utilize the text-mining tools used in Exercise 1 to analyze an observed association between epilepsy and autism spectrum disorders.
2.1 Using NER to dig deeper
The SCN2A gene is well known to be involved in epilepsy. To check if it has also been implicated in autism spectrum disorders, we will perform a systematic search for literature linking the two. A simple PubMed search retrieves only 14 publications:
https://www.ncbi.nlm.nih.gov/pubmed/?term=%22SCN2A%22+%22autism+spectrum+disorders%22
Since SCN2A (ENSP00000490107) and autism spectrum disorder (DOID:0060041) are both named entities in our dictionary, we can instead use the results of NER to retrieve relevant documents:
The NER-based approach retrieves many more publications. Inspect some of these abstracts.
12. Are they relevant and why were they were not found by the initial search in PubMed?
2.2 Linking diseases via genes
Above, we saw how existing text-mining resources can be used to retrieve abstracts connecting entities of interest and to extract associations. We will now use this in a different way to attempt to find genes that link two diseases of interest.
The first idea is to use NER-based information retrieval to find abstracts that mention both epilepsy (DOID:1826) and autism spectrum disorder (DOID:0060041). Click the link below to view these abstracts:
13. Which, if any, genes do you see mentioned in abstracts in the first two pages?
Note: The genes will not be highlighted with any color this time.
This approach obviously only works when the association between the two diseases is sufficiently well described in the literature for there to be abstracts that mention both diseases as well as the genes of interest. If that is not the case, but one has a candidate gene in mind, one can instead use information extraction to obtain a list of diseases for the gene in question to see if it is in fact associated with both diseases in the literature. Go to https://diseases.jensenlab.org/, query for SCN2A, and view the disease associations obtained from text mining.
14. Is SCN2A associated with both diseases of interest?
When one does not have a candidate gene, the best solution is to obtain two gene lists, one for each disease, and then identify the ones that rank high on both lists. This is a variant of the closed knowledge discovery problem. It is unfortunately not currently possible to perform such an analysis via a web interface, but it can be done by downloading the complete set disease–gene association from the DISEASES downloads page and analyzing them in Python or R. Alternatively, the analysis can be performed through Cytoscape as illustrated in the Cytoscape stringApp exercises.
Supporting Video Lectures
Supporting literature
Jensen LJ, Saric S and Bork P (2006). Literature mining for the biologist: from information retrieval to biological discovery. Nature Reviews Genetics, 7:119–129.
Abstract Full text
Fleuren WWM and Wynand Alkema W (2015). Application of text mining in the biomedical domain. Methods, 74:97–106.
Abstract Full text
Pafilis E, Buttigieg PL, Ferrell B, Pereira E, Schnetzer J, Arvanitidis C and Jensen LJ (2016). EXTRACT: Interactive extraction of environment metadata and term suggestion for metagenomic sample annotation. Database, 2016:baw005.
Abstract Full text
Grissa D, Junge A, Oprea TI and Jensen LJ (2022). DISEASES 2.0: a weekly updated database of disease–gene associations from text mining and data integration. Database, 2022:baac019.
PubMed Full text